We are very excited with the news that Evelina and Johan were both awarded an early-career support grant by the recently formed A-life. These grants will allow both of them to appoint a PhD student and to further develop their research endeavors through new collaborations within A-life.
Author: Johan van Heerden
In press: Rate maximisation makes the most complex microbes behave simple!
Our theoretical work on rate maximisation in metabolism has been published in Plos Computational Biology. In this work, we took a very general approach to ask a very specific question: we included genome-scale metabolic networks, complicated enzyme kinetics, and arbitrary constraints on enzyme expression, and then asked what metabolism looks like for a microorganism that maximises its biomass production. The answer: it looks quite simple. Only a few independent pathways are used by the microbe, giving rise to the linear relations and growth laws that are found across microbiology. With this result, we can not only model important phenomena such as overflow metabolism and co-consumption, but we can even understand them.
Over the past years, we have worked on this project a lot and we are very proud of the result. Daan: “Personally, I got the best introduction to academics and systems biology that I could have hoped for: from mathematical optimisation approaches (taught by Bob), doing lab work and coping with the corresponding disappointments (taught by Coco), to thinking, theorizing, writing and picking colours for your figures (taught by Bas and Frank).” We hope that our general, theoretical approach, will lead to more results and a better overall understanding in the near future.
The full open access article can be found here: https://doi.org/10.1371/journal.pcbi.1006858
In press: Dennis’ extensive characterisation of Fluorescent protein behaviour in yeast is published!
Dennis’ work on the
You can find the open access article here.
Two papers published by Lucas
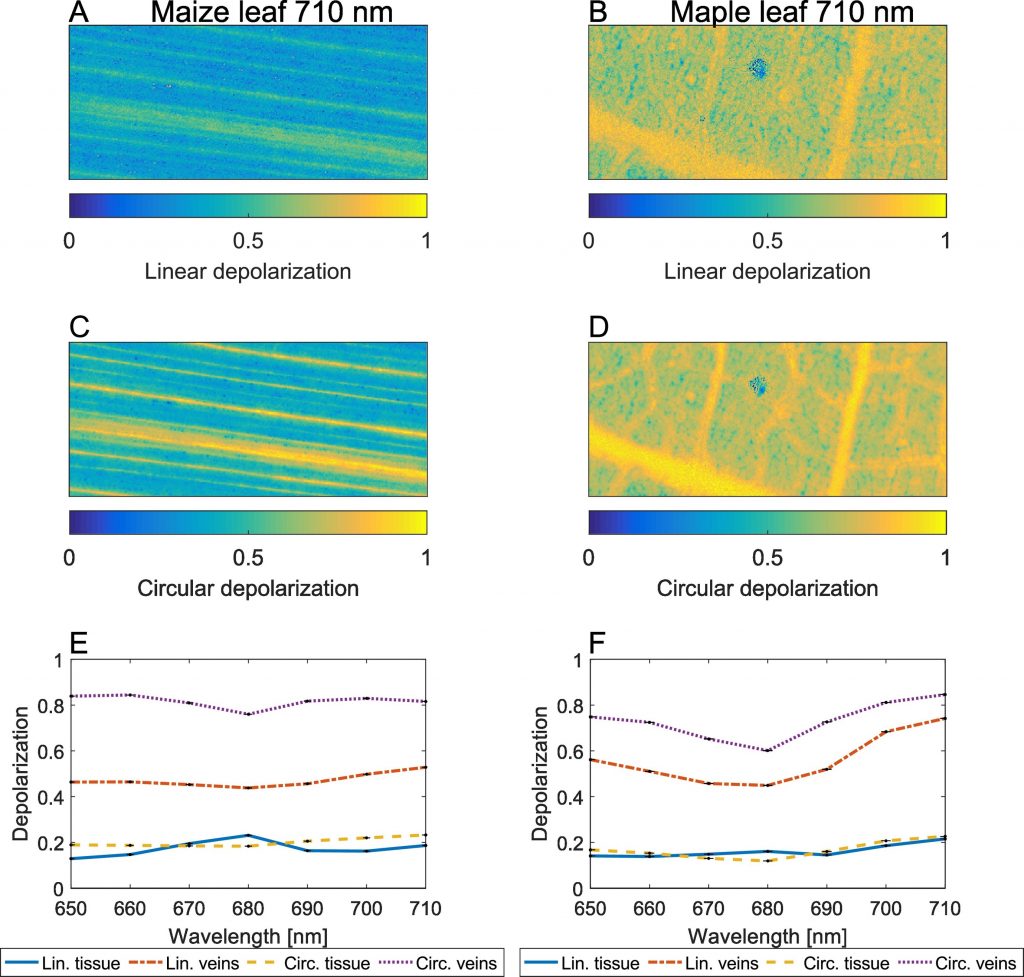
Lucas Patty has yet, after his recent publication in the Journal of Quantitative Spectroscopy & Radiative Transfer (dec 2017), published another paper! The new paper is entitled “Imaging linear and circular polarisation features in leaves with complete Mueller matrix polarimetry”.
Previously they showed that the photosynthetic apparatus in leaves induces circularisation ofunpolarised light due to its isotropic components, in the same amount as that it can absorb circularly polarised light. As these transmission measurements can be executed both in-vivo and post-mortem, they were able to confirmed that healthy leaves can successfully be differentiated from senescing leaves in-vivo.
In the new paper, Lucas shows that not only the photosynthetic apparatus, but also the cell structure and its arrangement within a leaf can create and modify polarisation signals. Basically, both linear and circular depolarisation is much larger in the veins than in the normal tissue.
Such in-dept understanding of polarisation spectroscopy data can both contribute to agriculture and the remote sensing of life on other planets!
Niclas Nordholt got his PhD
Niclas defended his PhD thesis, titled “Balance on many scales: Growth and gene expression in Bacillus subtilis” on January the 18th 2018. James Locke, Frank Schreiber, Rutger Hermsen, Sander Tans, Leendert Hamoen and Bas Teusink were in the thesis committee. A big thanks to the committee for their efforts, and congrats to you Dr. Niclas!

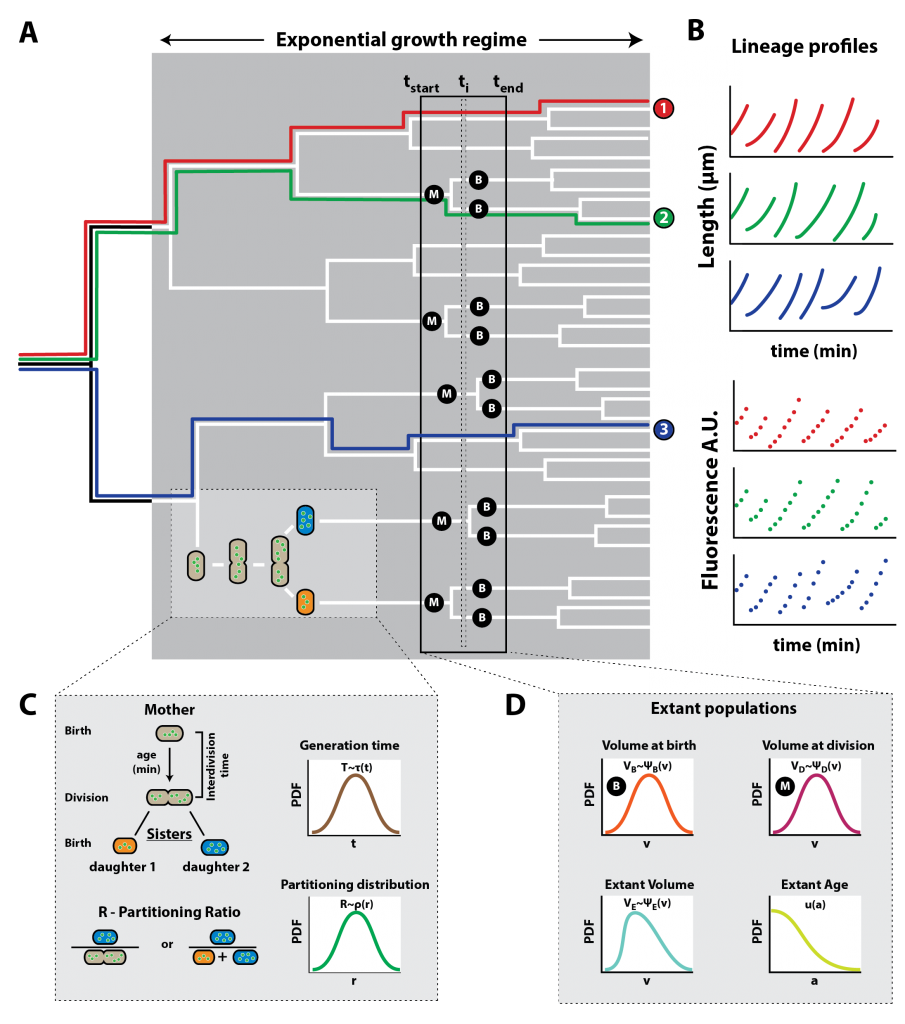
Our paper on the statistics of single cell bacterial growth is published
In this paper we describe a methodology for the prediction, analysis and tests of consistency of single-cell growth and protein expression data from a few
basic statistical principles.
You can find it here.
Johan van Heerden got his PhD
On January 11th Johan van Heerden defended his PhD entitled: “Dynamic regulation of yeast glycolysis through trehalose cycling: a probabilistic view of metabolic transitions”, of which Bas was the promoter. The committee comprised of prof Stefan Hohmann, prof Hans Westerhoff, prof Barbara Bakker, prof Frank Bruggeman and Dr Aljoscha Wahl. Previous group members, Jan Berkhout and Filipe Santos were the paranymphs.
Dan Fraenkel and Jens Nielsen have written an in-depth review of our Science paper
Dan Fraenkel and Jens Nielsen recently undertook an in-depth review and extended analysis of our Science paper (van Heerden et al., 2014), and we are excited to see that their story has been published!
Fraenkel and Nielsen, 2016. Trehalose-6-phosphate synthase and stabilization of yeast glycolysis
Our time-lapse segmentation and tracking algorithm can now also track growing S. pombe
We have been hard at work developing an automated segmentation and tracking tool to analyse microscopic time-lapse movies of growing bacteria. After a few minor adjustments and additions we can now also segment and track fission yeast (Schizosaccharomyces pombe) growing on an agarose pad.
Our algorithm is now in its final stages of development and testing, and should be ready for release early next year.

Recent Comments