Single-cell Physiology
Below, you can find the updated list of Bachelor and Master internships available at the moment.
For more information on available internships in the Single-cell Physiology Team, please contact directly the team leaders:
Evelina Tutucci – evelina.tutucci@vu.nl
Johan van Heerden – j.van.heerden@vu.nl
| Project title | Type of research | Supervisor(s) |
|---|---|---|
| Developing smRNA FISH for the identification of antibiotics-tolerant persister bacteria | Experimental (Master) | Evelina Tutucci (evelina.tutucci[at]vu.nl) Wilber Bitter (w.bitter@vu.nl) Frank Bruggeman (f.j.bruggeman@vu.nl) |
| Motivation Multi-drug-resistant, pathogenic bacteria threaten human health. They have acquired DNA mutations that make them insensitive to antibiotics. Pathogenic bacteria are also successful in dealing with antibiotics, and the human immune system, because they often can switch to a non-growing, antibiotics-tolerance state, called the ‘persister’ state. Persister cells are formed from growing bacteria, can switch back to the growing state and coexist with growing cells in cell cultures. Since persister cells are not growing, they can withstand the effects of antibiotics that kill growing cells. It has been shown that growing cells can switch into persister cells, via so-called toxin-antitoxin systems. Such systems have been discovered in Escherichia coli and have since been found in many pathogenic bacteria, including Pseudomonas aerogenosa and Mycobacterium tuberculosis. Understanding how persister cells form is an important research topic. Aim of this project Since persister cells derive from growing cells and coexist with them in growing cell culture, we require single-cell methods to identify persister cells. One promising method is to determine the relative number of toxin and antitoxin transcripts in single cells, as it is believed that an excess of toxins over antitoxins turn growing cells into persister cells. The aim of this project is develop single-molecule RNA FISH, which allows for counting of the number of transcripts in single cells, for identification of persister cells. First in Escherichia coli and subsequently in Mycobacterium segmatus, a relative of M. tuberculosis. Methods and techniques fluorescence microscopy, molecular biology, bacterial cell cultivation, RNA methods, smRNA FISH. Impact During this project you become familiar with state-of-the-art methods of microbiology, molecular biology and fluorescence microscopy that are used throughout molecular biology labs in hospitals, academia and industry. Since smRNA FISH is not routinely used in such labs, the expertise which you acquire during this project will count on the job market, regardless whether this concerns a fundamental or applied job. Number of positions available 1 Duration of project 6 months Starting date As soon as possible | FILLED until September 2024 | |
| Developing and using a new cyan-based glucose sensor | Experimental (Bachelor or Master) | Dennis Botman d.botman@vu.nl |
| Glucose is the major carbon source for Saccharomyces cerevisiae to grow on. In order to measure glucose levels in single living yeast cells, biosensors based on fluorescent protein are needed. Currently, none of the fluorescent biosensors for glucose are suitable as they are dim or pH-sensitive. We have developed 2 new glucose biosensors based on a cyan fluorescent protein and further development and characterization is needed. Once these sensors prove to be robust tools to measure intracellular glucose levels, we will use it to elucidate glucose metabolism in budding yeast. | ||
| Filamentous Growth and Biofilm formation in S. cerevisiae and C. albicans resolved by single cell imaging | Experimental (Master) | Evelina Tutucci (evelina.tutucci@vu.nl) |
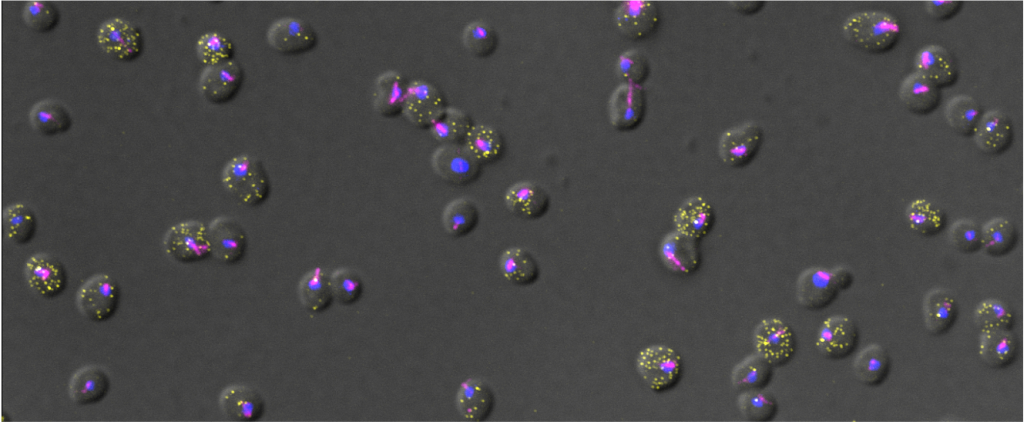
| Background Many fungi such as Saccharomyces cerevisiae or Candida albicans are able to switch between a unicellular (yeast) form to a multicellular filamentous form in response to changes in the environment (e.g. nutrients availability, stress). This morphological transition allows fungi to adopt different survival strategies and in some instances become pathogenic. During filamentous growth cells acquire an elongated shape and unipolar budding pattern, allowing for greater exploration of the environment. A more advanced strategy is the formation of biofilms, multicellular structures that consist of different cell types (both yeast form and filamentous form) as well as an extracellular matrix, offering both increased structural integrity and resistance to antifungal drugs. While many of the genes required for this differentiation process have been identified through bulk analysis (e.g. RNA seq), their expression in single cells and during differentiation has remained largely unstudied. Aim In this project, we investigate at the single cell level, the gene expression changes occurring during fungal differentiation. By using a fluorescence-based RNA imaging technique called smFISH (see pictures: https://www.tutuccilab.com/) we visualize and quantify individual mRNA molecules in single cells to investigate the spatiotemporal control of gene expression during filamentation. Furthermore, we investigate how the spatial organization of cells in biofilms influences gene expression. Planned activities (and methods) During this project, you will learn how to cultivate filamentous fungi, various molecular biology techniques as well as smFISH and cutting edge fluorescence microscopy approaches. You will also be trained in imaging analysis, which will be used to investigate gene expression (RNA spot counts) of cells in biofilm. We will measure cell-to-cell heterogeneity and couple it with spatial information, cell volume and cell length in filamentous cells. Since the microscopy data is very rich of information it will be possible to further expand the analysis, depending on your computational skills and your curiosity. A prior familiarity with programming languages such as Python and R is a must . You will participate and present in our Single-cell group meeting and Journal club. Number of positions available 2 Duration 6 Months Starting date As soon as possible | Available | |
| Decisionmaking in single cells during nutrient shifts | Experimental (Master) | Philipp Savakis p.e.savakis[at]vu.nl |
| Background Microorganisms have to make decisions continuously. In environments with changing nutrients, these decisions can include a complete reversal of the direction of metabolism. This is inherently risky: cells that switch too early miss out on favourable conditions, while late adapters may be too late to the party. Whole cultures can employ bet hedging strategies to offset the risk of incorrect decision making, but it is not clear how individual cells contribute to this emergent behaviour. Aim In this study, we investigate the decisionmaking in single cells of the yeast Saccharomyces cerevisiae during nutrient transitions, using a combination of live cell imaging as well as fixed cell techniques. For the latter, we will pinpoint the moment at which single cells commit to metabolic change by visualising single mRNAs using fluorescence-based single molecule techniques. In the live-cell imaging part we will subject mutant strains with fluorescently labeled metabolic enzymes directly to nutrient transitions using microfluidic cultivation. What you will do - Construct mutant S. cerevisiae strains and characterize their behaviour using microfluidics and time-lapse microscopy. - Visualise single mRNAs using smFISH - Learn to use a variety of image processing tools based on neural networks Duration 6 months Starting date sept/oct 2023 |
||

Recent Comments